You’ll probably remember this one from grade school biology. Mitochondria are ‘the powerhouse of the cell’. Which basically means that mitochondria take sugars and break them down to make energy.
But those little powerhouses were once so much more- they were an entire organisms. A very long time ago, a certain type of proteobacteria was engulfed by a single celled host organism, but that organism decided that instead of digesting the proteobacteria, it would hold onto it. With time, the proteobacteria evolved into the modern mitochondria, a process that involved most of its very own bacterial genome being stolen away, and sequestered in the nucleus of the host.
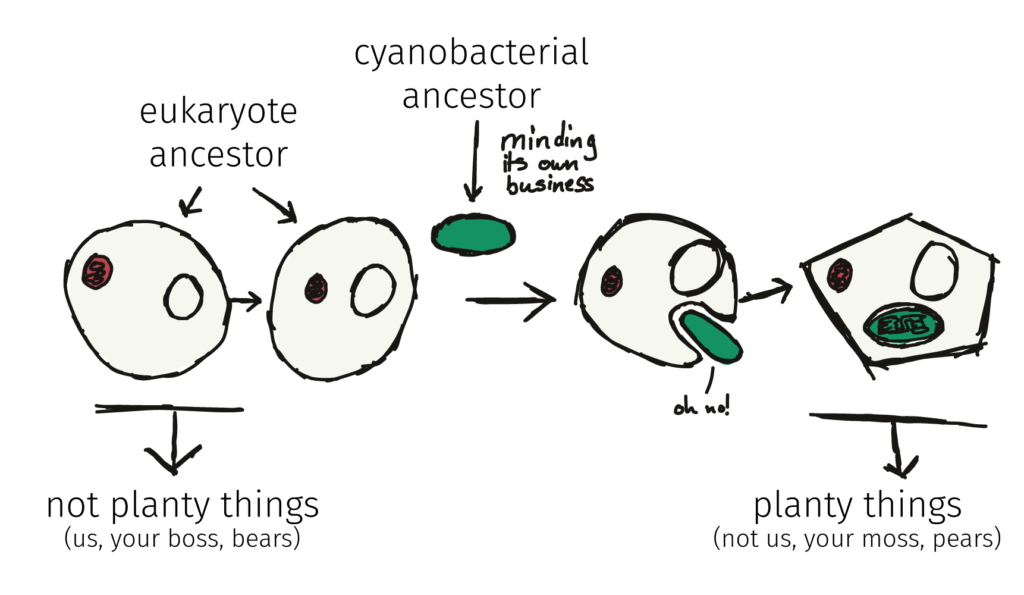
By the way, a similar engulfment, this time of a cyanobacteria, initiated the development of the chloroplast, and ultimately, the birth of ‘plants’. We’ve written about that one over here.
Nowadays, the plant mitochondrial genome only encodes approximately 20 proteins, but the plant mitochondria needs about 2000 proteins to function (i.e., to be all powerhouse-y). Expressing a whole lot of proteins from the nucleus, and then having to both tell the proteins where to go and actually get them into the mitochondria where they belong, might seem like a lot of extra effort. But it allows the nucleus to control exactly what happens and when.

Don’t forget, the nucleus is the brain of the cell!
So fairly unsurprisingly, the majority of those 2000-or-so nuclear-encoded powerhouse-y proteins look like they were originally stolen (transferred) from the mitochondrial genome- we can see that the genes encoding them look pretty proteobacterial. But, every now and then, we see something that is used in the mitochondria, but looks like a non-stolen nuclear gene.
So how do plants make new proteins to control their ex-proteobacterial mitochondria? Well sometimes, it seems, they plagiarize from themselves.
A recent study (Lama et al.) investigated a family of proteins known as DUF295 (DUF stands for ‘Domain of Unknown Function’, if that helps at all). The gene family isn’t found in animals, and even in plants, seems to be pretty much constrained to the angiosperm (flowering plant) group. In fact, some types of the DUF295 family seem to only be in the found in the Brassicacae family, and our favourite Brassicaceae Arabidopsis has NINETY FOUR different DUF295 genes. This recent evolution of huge numbers of genes, and the physical location of these genes together on the chromosomes, suggest that many DUF295s have been created by simply copying chunks of genome, and pasting it in another location.

Despite the obvious ‘copy-paste’ function, there’s evidence that these different DUFs are starting to develop their own personality. Many of them are expressed under fairly specific conditions, either in specific plant tissues (leaf, pollen, seeds), at certain times, or in response to abiotic or biotic stresses. Furthermore, while some of the genes seem to be lowly expressed and even missing from certain accessions (races) of Arabidopsis, others seemed to be conserved, suggesting that they might be serving a useful function and therefore need to be actively kept functional.
We’ve written about Arabidopsis accessions, and how they can be a powerful tool for plant research here.
Amazingly, some of the new DUF295 proteins seem to head to the mitochondria- as shown by experiments that hitch a visible fluorescent marker (GFP) to the back of certain DUF295s. In fact, many of the newly-created DUF295 genes have lost a certain functional domain (F-box domain) usually found at the start of their protein, and replaced it with a short ‘targeting sequence’ that guides the protein product into the mitochondria.

Once in the mitochondria, it looks like at least some DUF295s are involved in retrograde signalling, a process that involves the mitochondria communicating its status, and its needs, back to the nucleus. Weirdly, this signalling pathway involves proteins whose genes have been around for much longer than the mitochondrial DUF295s, suggesting that these new kids on the block have somehow managed to insert themselves into an already-existing pathway.
As a final experiment to further figure out the function of the mitochondrial DUF295s, the authors did what all good scientists do: they got mutated plants that didn’t accumulate individual DUF295s (T-DNA insertion lines) and looked to see if the plants were sick.
Perhaps surprisingly, given how important that mito is in being a cellular powerhouse, the plants weren’t troubled at all. Which might just go to show, that the copy-past function of gene duplication might not only allow the development of new functions, it can also give a little bit of redundancy to protect plants from prying scientists!
References
This post is based on the paper:
Lama S, Broda M, Abbas Z, Vaneechoutte D, Belt K, Sall T, Vanderpoele K, Van Aken O. 2019 Neofunctionalization of Mitochondrial Proteins and Incorporation into Signaling Networks in Plants. Mol Biol Evol 36(5): p. 974-989.
The role of DUF295s was suggested on the basis of previous research, which showed induction of eight DUF295s in mitochondrial mutants:
Van Aken O, Ford E, Lister R, Huang S, Millar AH. 2016. Retrograde signalling caused by heritable mitochondrial dysfunction is partially mediated by ANAC017 and improves plant performance. Plant J. 88(4): 542–558.
